Proteins are dynamic molecular machines whose function depends on coordinated conformational motion and interdomain communication. When these dynamic processes are disrupted, the consequences range from catalytic inefficiency to disease progression and therapeutic resistance. Our laboratory integrates protein engineering and quantitative biophysics to define how conformational landscapes control function, and to translate these insights into engineered catalytic systems and mechanistic intervention strategies across human health, pharmaceutical, chemical, and sustainability applications.
Catalytic Protein Discovery & Engineering
We discover and redesign catalytic proteins to expand the chemical capabilities of biology. Our work spans enzymatic fluorination, carbon capture enzymes, and new-to-nature chemistry, with an emphasis on mechanistic understanding and rational redesign.
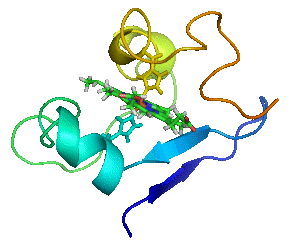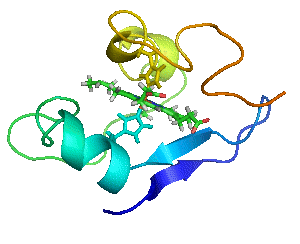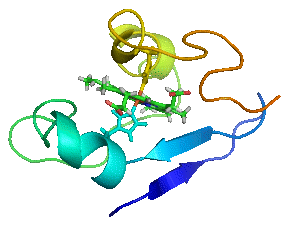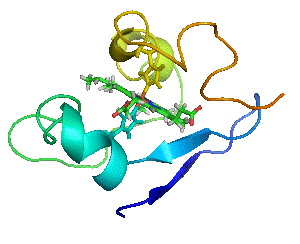
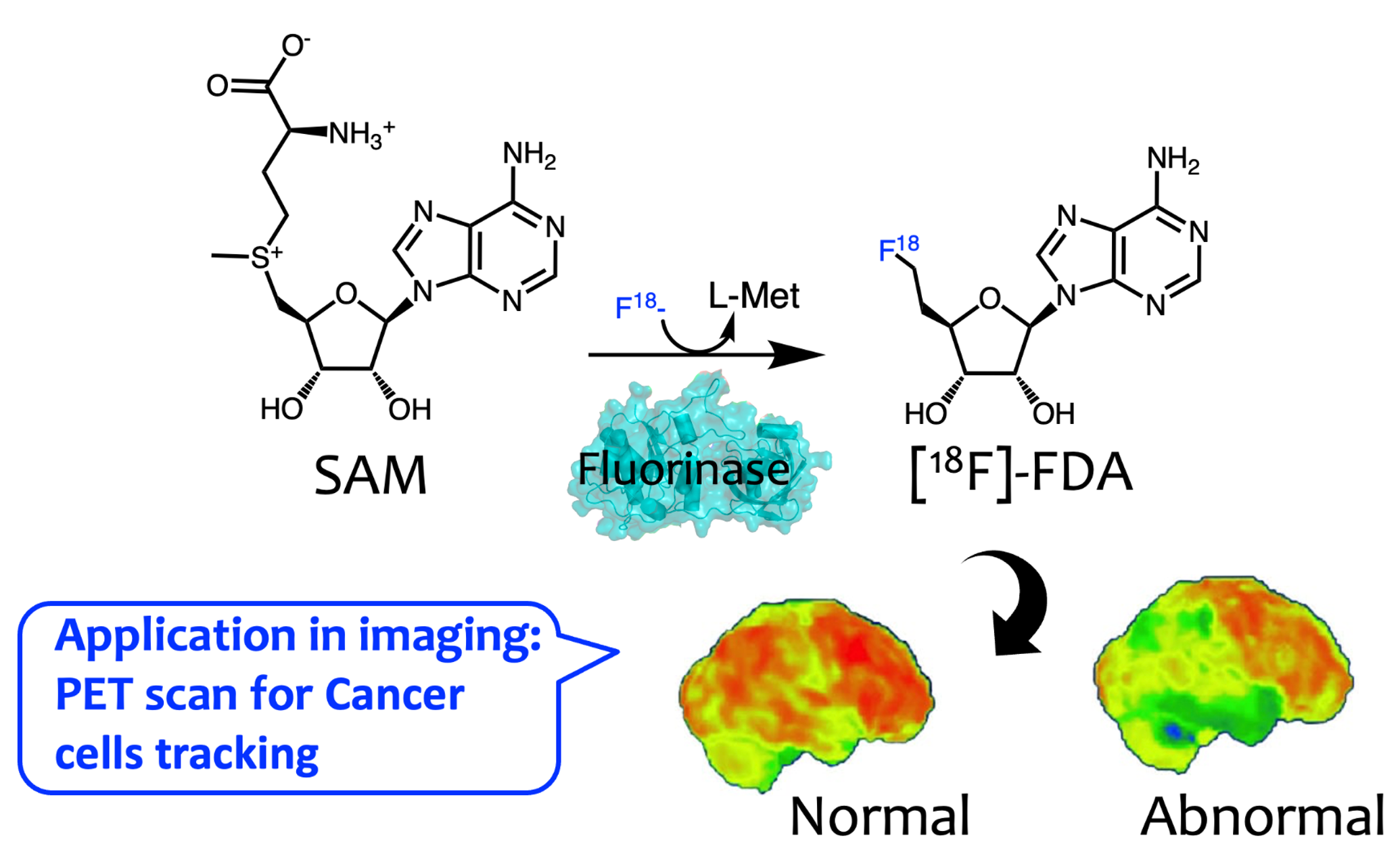
🔹 Fluorinase Discovery & Engineering
We identify and characterize new fluorinases to expand biological fluorination chemistry. Through structure-guided mutagenesis and kinetic analysis, we define active-site and long range determinants governing substrate scope and catalytic efficiency. These studies enable redesign of enzymatic C–F bond formation for radiochemical applications
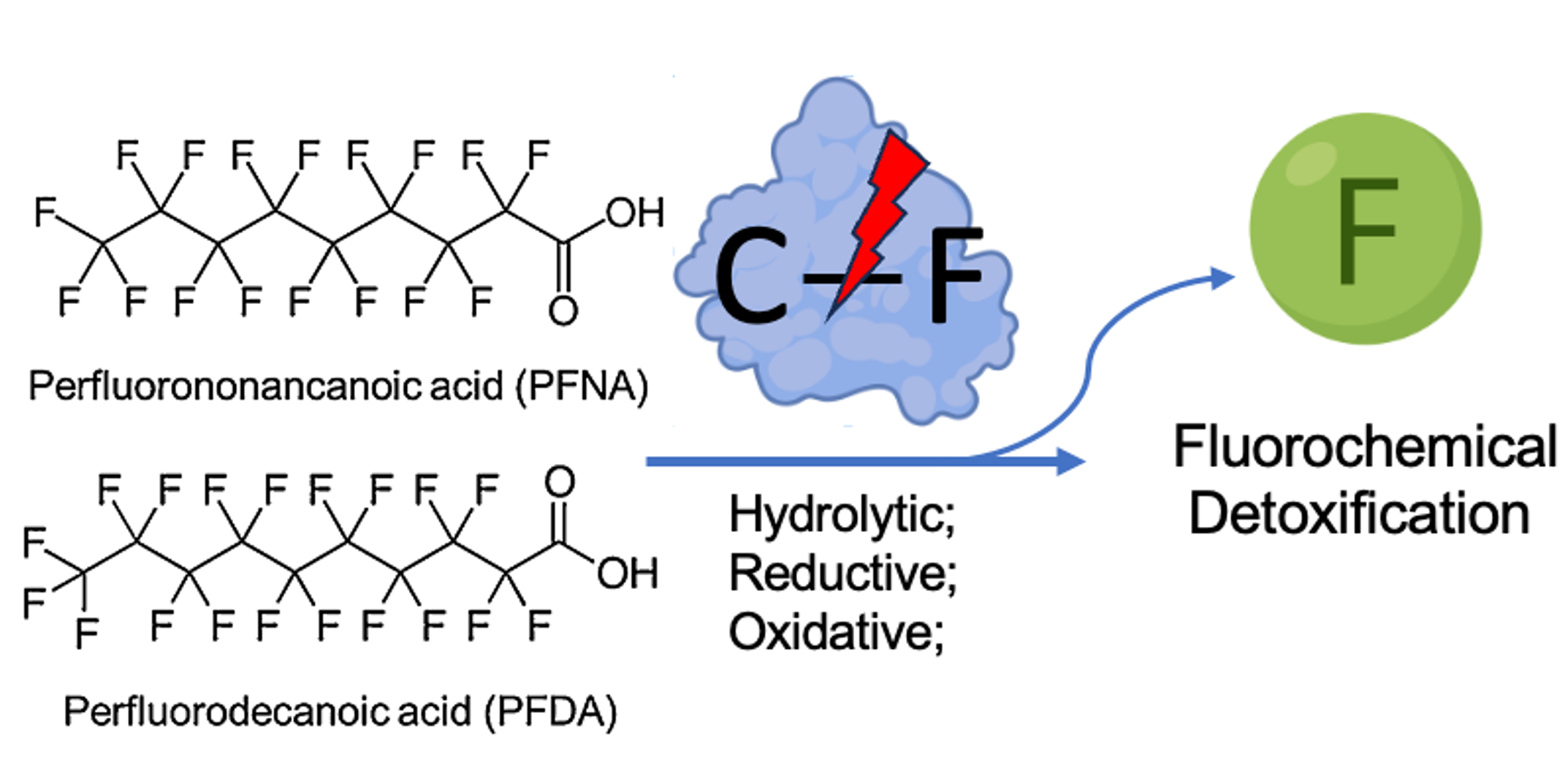
🔹 Defluorinase Engineering for Detoxification
To address the environmental persistence of fluorinated compounds, we investigate enzymatic strategies for C–F bond cleavage. By mapping catalytic mechanisms and engineering active-site architecture, we aim to develop biocatalytic platforms for fluorinated compound transformation.
🔹 Carbonic Anhydrase: Mechanism & Environmental Applications
Carbonic anhydrase provides a model system for understanding zinc-dependent catalysis and proton transfer networks. We combine mechanistic enzymology with protein engineering to explore enhanced CO₂ capture and environmental applications, while dissecting structure–function relationships in metalloenzymes.
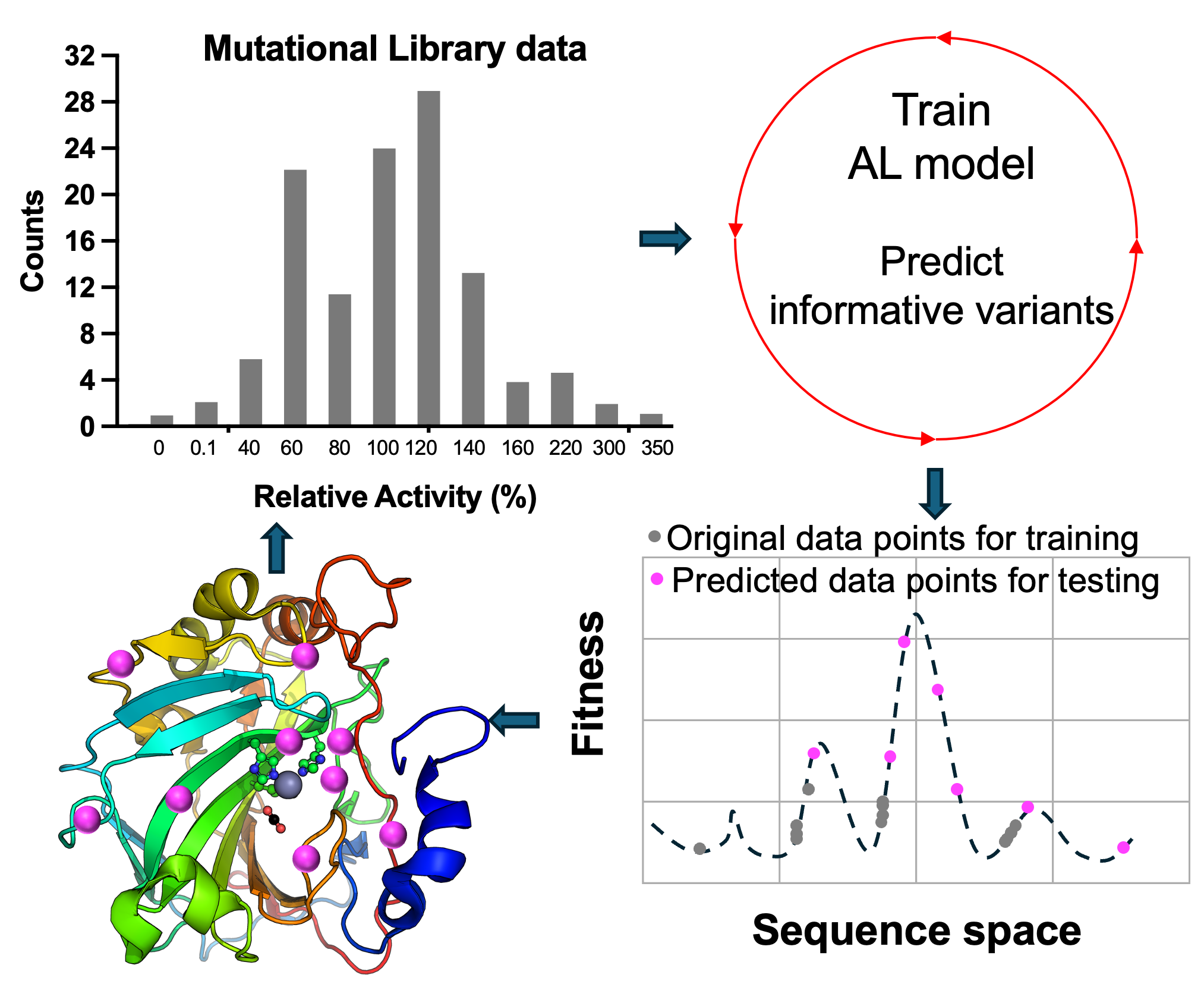
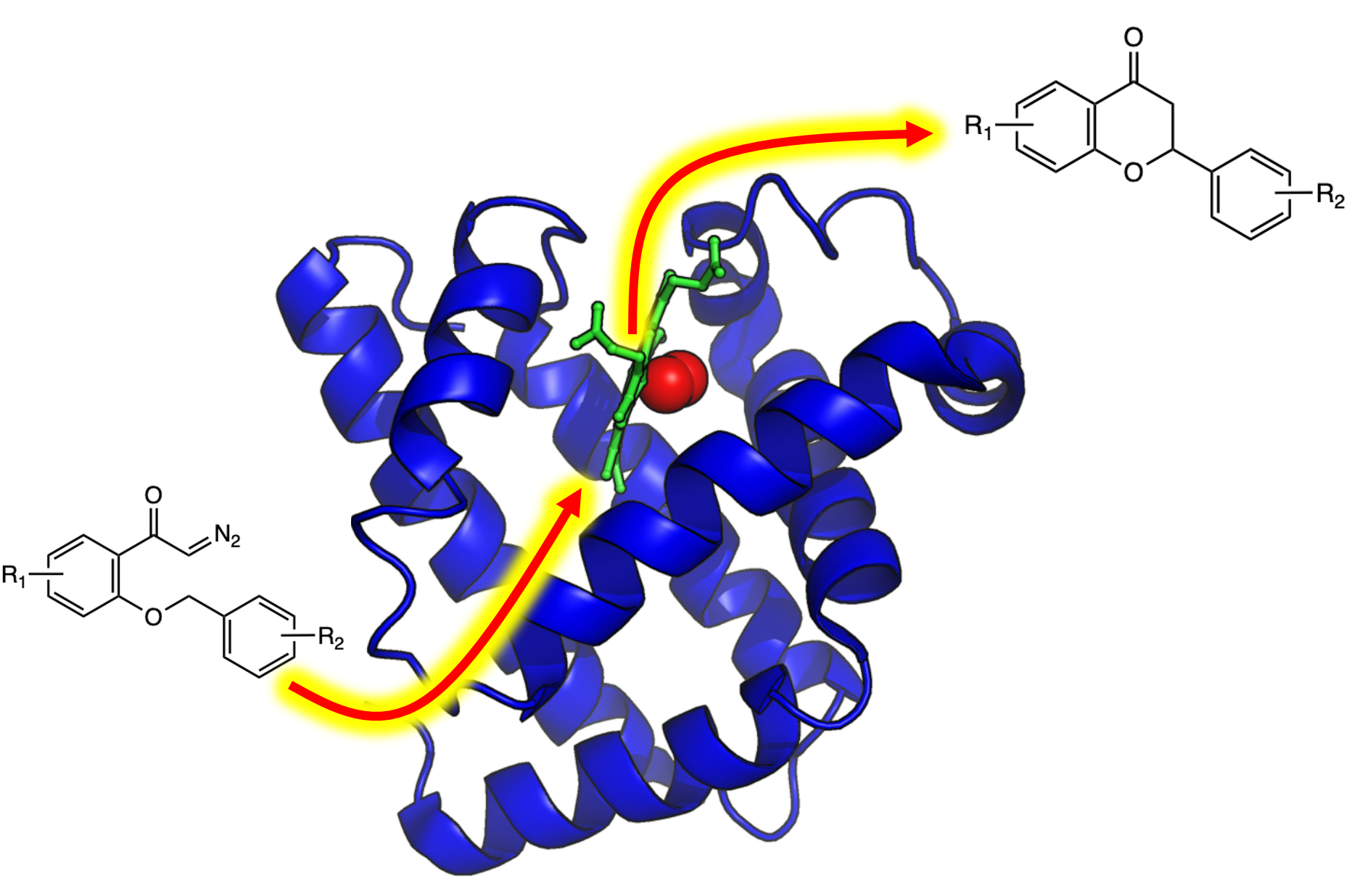
🔹 Myoglobin as a Model for Protein Dynamics
Myoglobin provides a well-defined model system to interrogate conformational exchange and folding dynamics using high-resolution NMR spectroscopy. By quantifying how structural fluctuations shape stability and functional plasticity, we uncover biophysical principles that govern evolvability and adaptive landscapes. These insights inform our efforts to engineer proteins capable of supporting new-to-nature chemistry by revealing how dynamic features enable the emergence and optimization of novel catalytic functions.
Biophysics of Disease-Related Proteins
Protein function is governed by dynamic conformational landscapes. We use HDX-MS, NMR spectroscopy, mutagenesis, and computational modeling to define how mutations, ligands, and post-translational modifications reshape structural communication in disease-relevant systems.
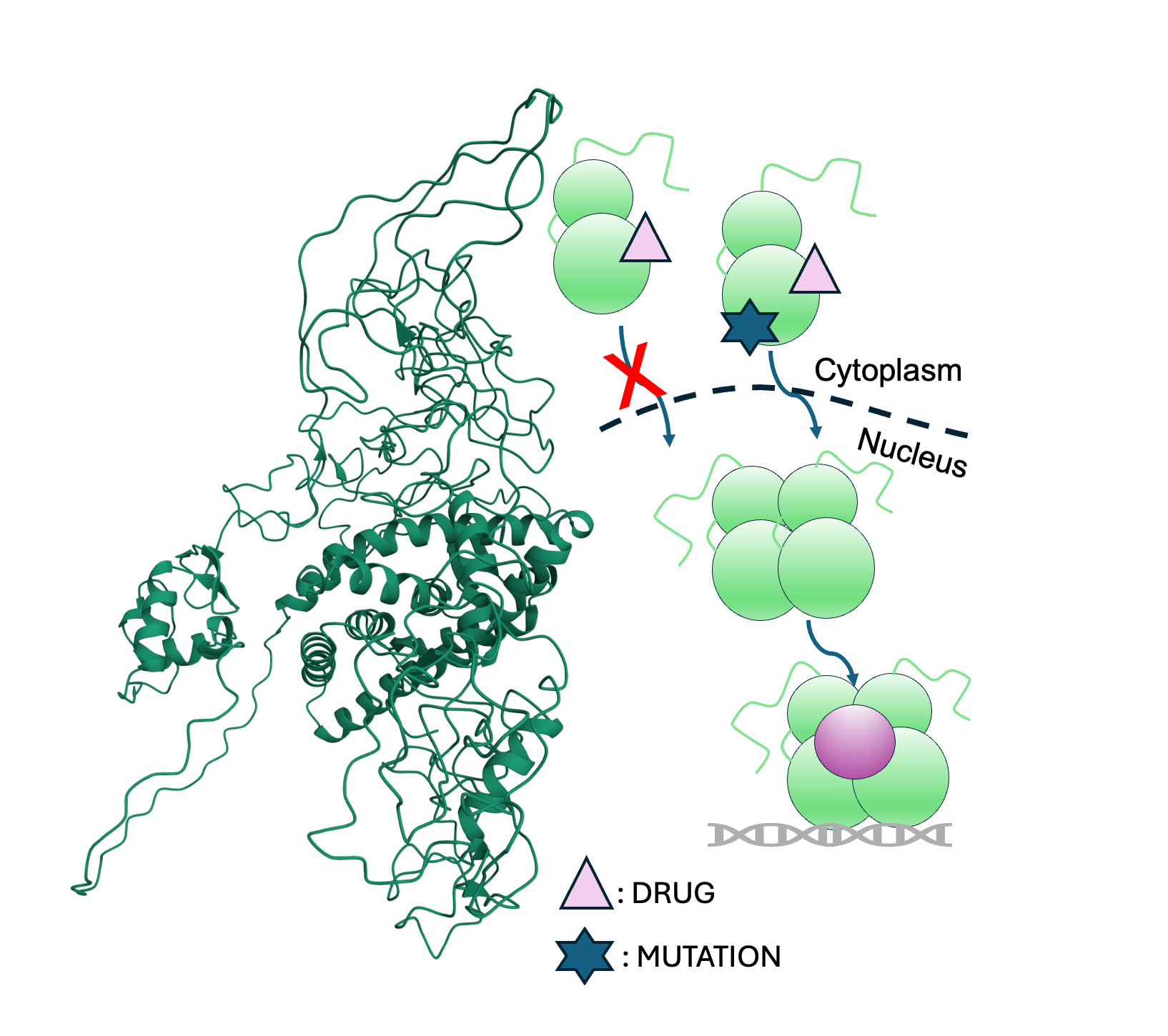
🔹 Estrogen Receptor α (ERα)
Estrogen receptor α drives the majority of hormone-responsive breast cancers, yet resistance to endocrine therapies remains a major clinical challenge. We investigate how ligand binding, phosphorylation, and clinically observed mutations reprogram interdomain communication in full-length ERα. Using time- and temperature-resolved HDX-MS integrated with computational modeling, we define dynamic vulnerabilities that underlie aberrant signaling and therapeutic resistance. Our goal is to translate conformational insight into new strategies for selectively modulating ERα activity in resistant disease.
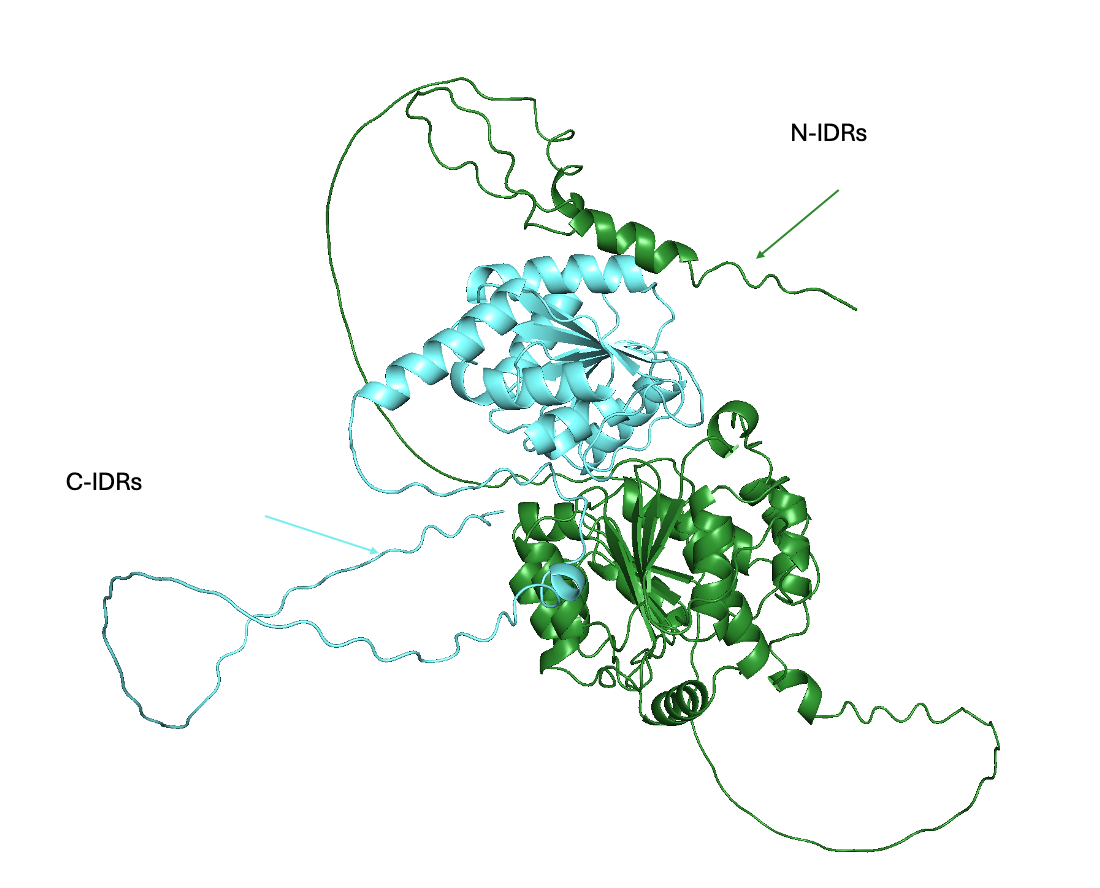
🔹 DDX3X RNA Helicase
DDX3X is an RNA helicase implicated in cancer, neurodevelopmental disorders, and antiviral signaling. Disease-associated mutations often disrupt its tightly coordinated conformational transitions, leading to altered RNA processing and dysregulated cellular pathways. We investigate how these mutations perturb dynamic coupling between domains and rewire functional output. By integrating HDX-MS, mutagenesis, and structural modeling, we aim to identify mechanistic control points that can be leveraged for therapeutic modulation.
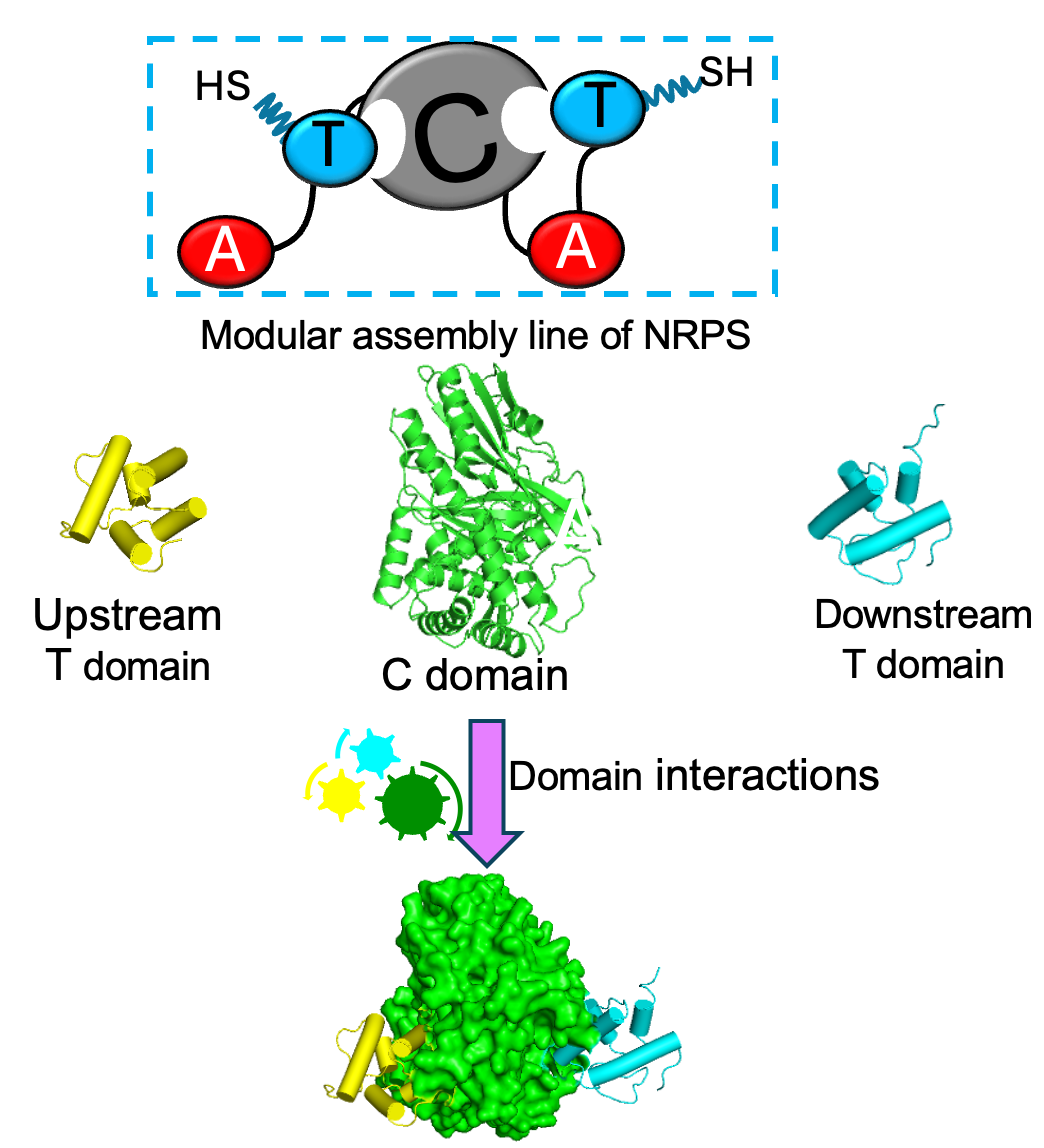
🔹 NRPS Systems & Modular Enzyme Interfaces
Nonribosomal peptide synthetases (NRPS) are dynamic, multi-domain assembly lines responsible for the biosynthesis of many clinically important natural products, including antibiotics and anticancer agents. We investigate how domain interfaces and conformational transitions coordinate catalysis across modules and how disruption of this communication limits biosynthetic efficiency and fidelity. By defining the dynamic principles that govern interdomain coupling, we aim to enable rational reprogramming of modular biosynthetic systems for the development of next-generation therapeutic compounds.