Protein Dynamics-Function Relationship
Why do we care about protein dynamics?
Proteins are not statues.
They breathe, fluctuate, and transition between conformational states.
Function is encoded not in a single structure, but in an ensemble of dynamically interconverting states. These motions determine how enzymes catalyze reactions, how receptors signal, and how mutations rewire regulatory networks.
Most drugs are designed against static structures — yet disease often emerges from shifts in conformational landscapes.
By mapping these dynamic ensembles, we uncover hidden regulatory states, identify conformational vulnerabilities, and define principles for engineering new catalytic function.
To control biology — whether to redesign enzymes or intervene in disease — we must first understand its motion.
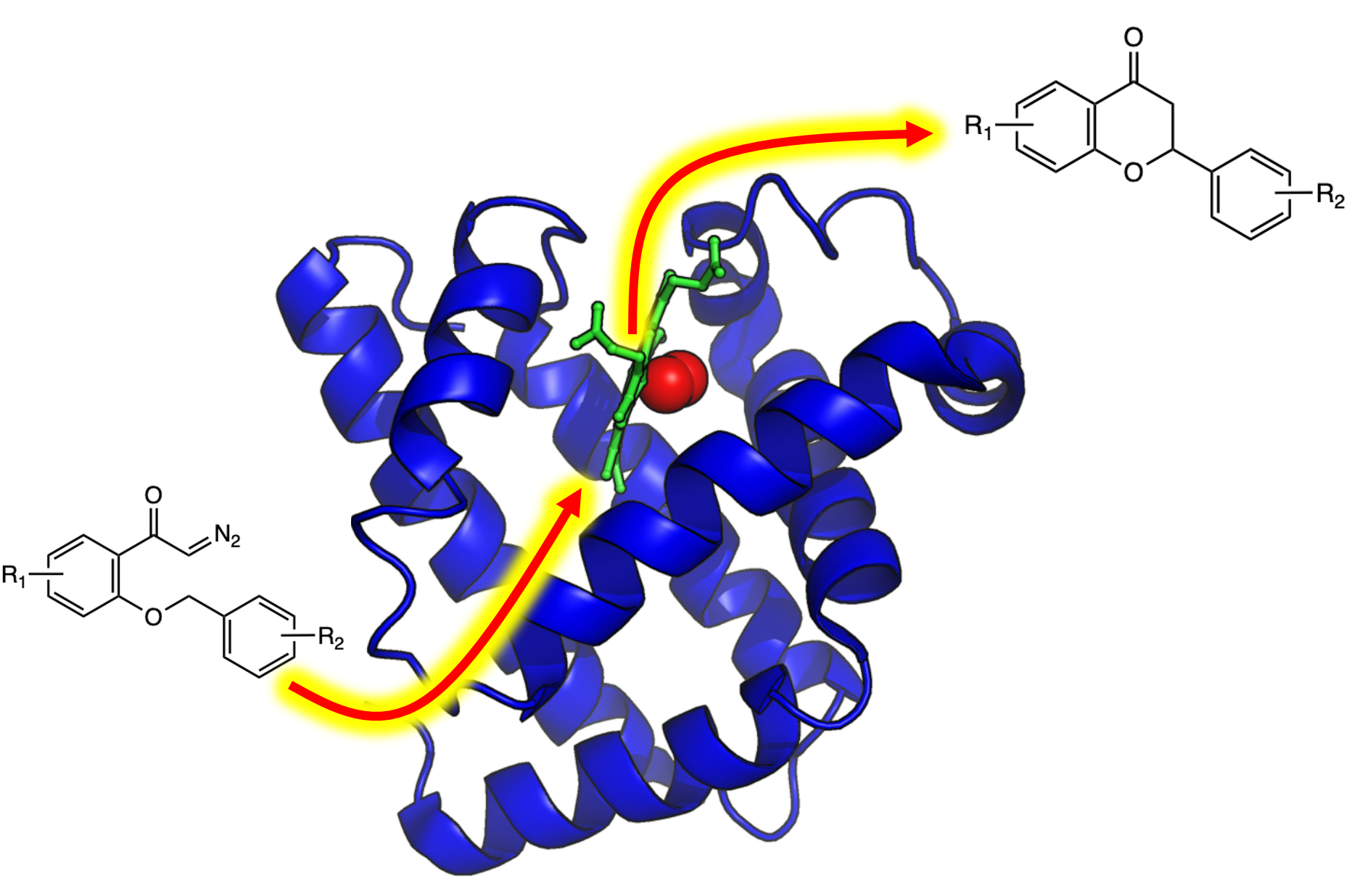
Myoglobin as a Model for Protein Dynamics
Myoglobin provides a well-defined model system to interrogate conformational exchange and folding dynamics using high-resolution NMR spectroscopy. By quantifying how structural fluctuations shape stability and functional plasticity, we uncover biophysical principles that govern evolvability and adaptive landscapes. These insights inform our efforts to engineer proteins capable of supporting new-to-nature chemistry by revealing how dynamic features enable the emergence and optimization of novel catalytic functions.
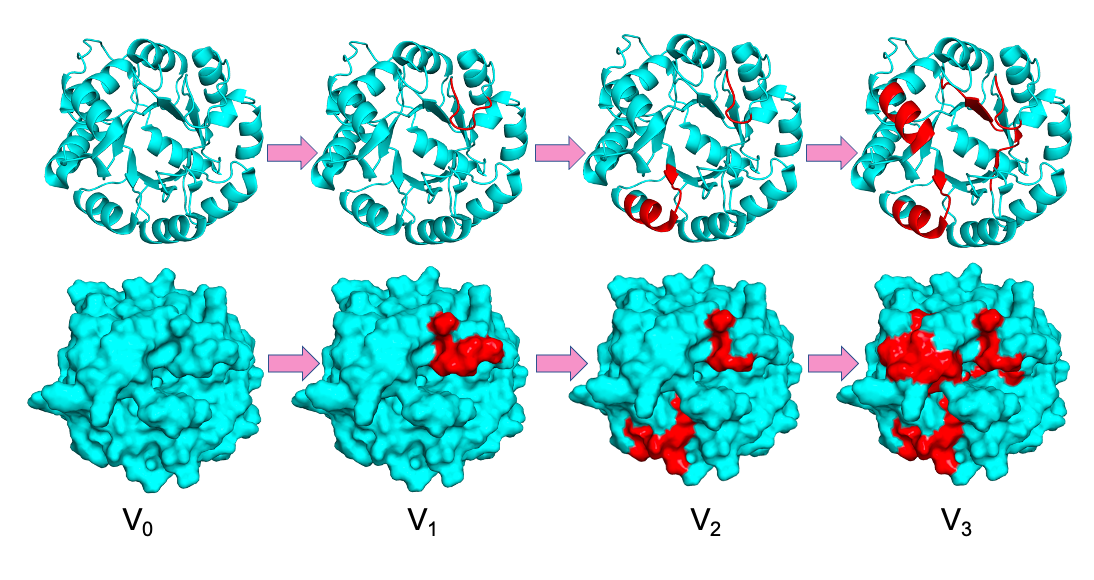
Investigating Protein Dynamics Along an Evolutionary Trajectory
We use time-dependent HDX-MS to map how conformational dynamics evolve across successive generations of engineered aldolase variants with progressively enhanced activity. By identifying regions where flexibility is selectively tuned, we uncover dynamic networks that enable optimized catalysis. These insights provide a framework for HDX-guided enzyme design and rational engineering of improved catalytic systems.